I am a Research Scientist at the Chan Zuckerberg Initiative AI/ML team.
I have completed my PhD under the supervision of Fabian Theis at the Computational Health Center, Helmholtz Munich and the Munich School for Data Science. Additionally, I am co-founder and core developer of scverse, an open-source ecosystem for single-cell data analysis.
vitae / scholar / github / linkedin
I am interested in AI and Software for Computational Biology, endurance sports and books.
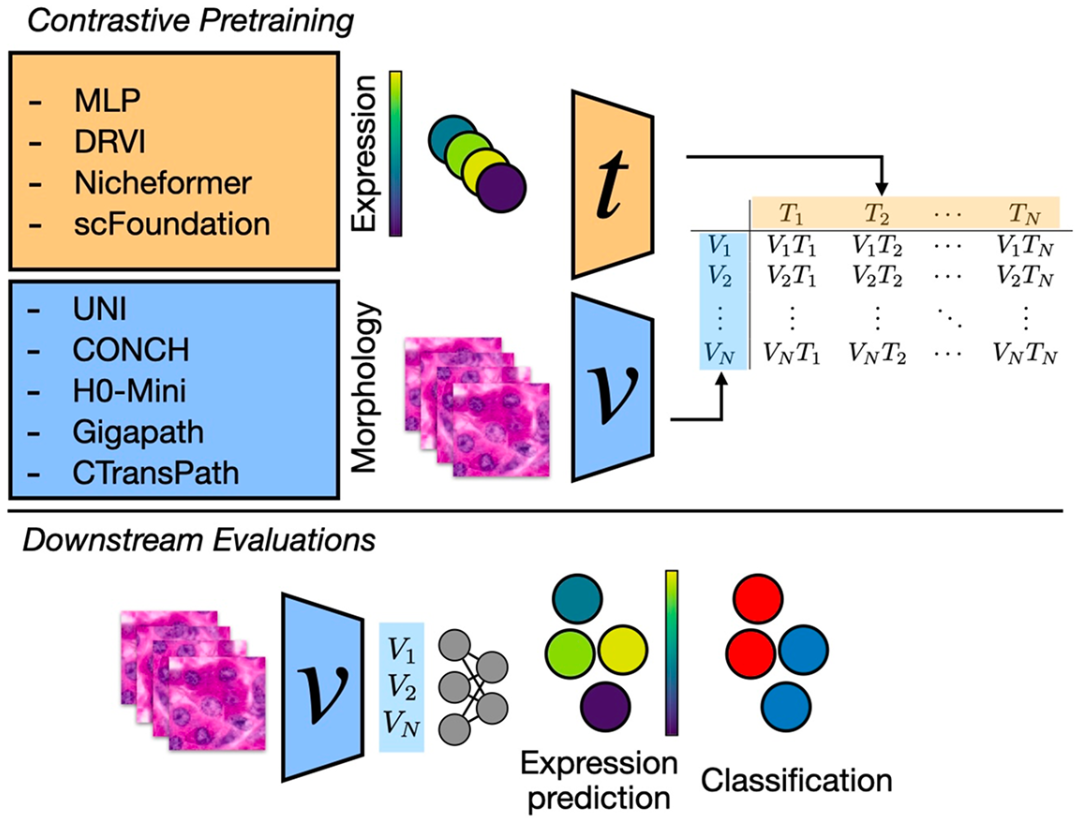
Hescape
A large scale benchmark for cross-modal learning of histology and spatial expressions.
Project Page | Paper (ICCV - CVAMD, 2025)

Moscot
A framework for mapping cells in time and space with optimal transport.
Project Page | Paper (Nature, 2025)
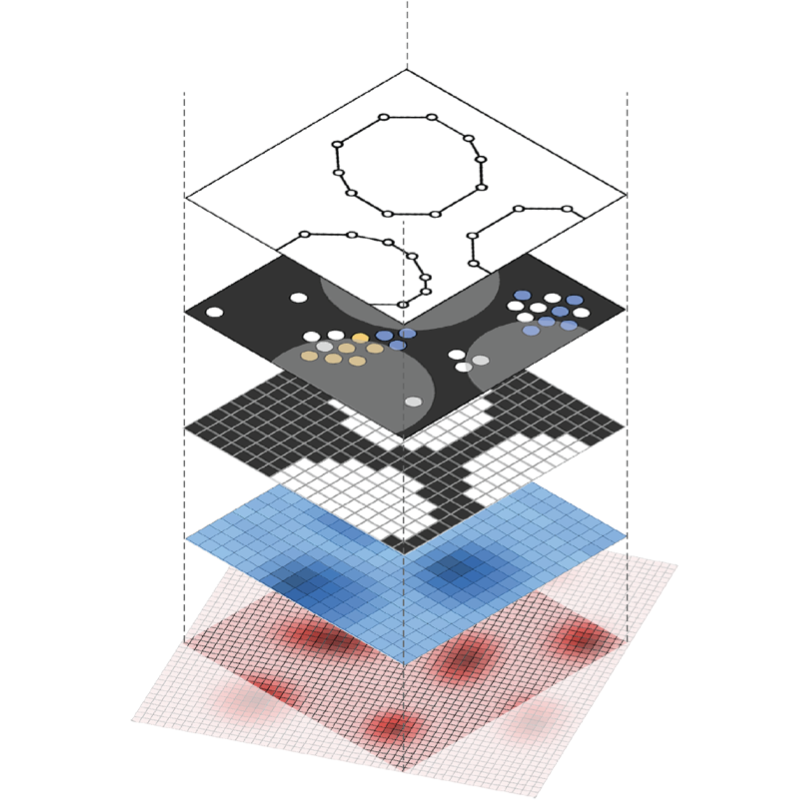
SpatialData
An open and universal framework for multi-modal spatial omics data.
Project Page | Paper (Nature Methods, 2024)

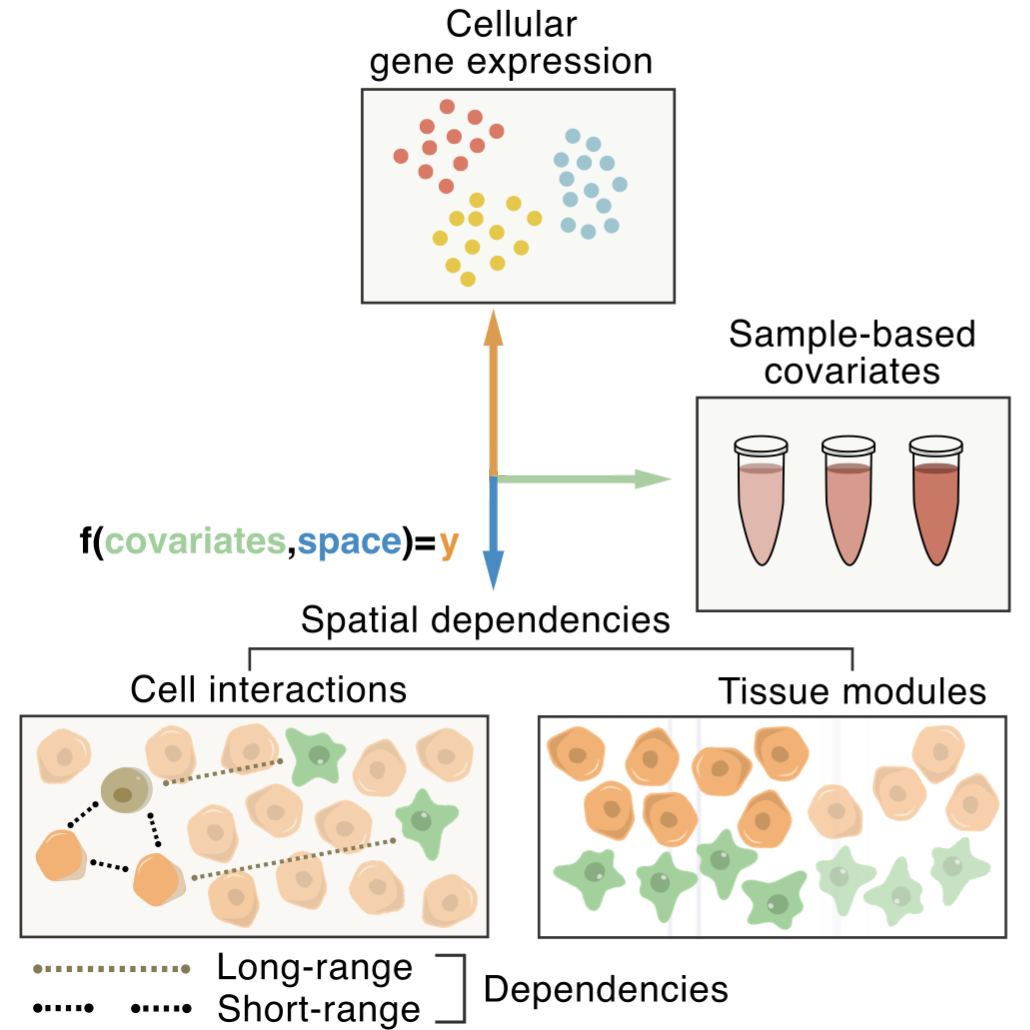
Spatial Components of Tissue Variation
A perspective and review on open problems in tissue biology.
Paper (Nature Biotechnology, 2022)
News
- [September 2025] I am co-organizing the 2nd scverse conference at Stanford. I will be chairing the panel on Agentic Workflows in Bioinformatics.
- [April 2025] I am co-organizing the workshop Learning Meaningful Representation of Life at ICLR 2025 in Singapore.
- [September 2024] I have joined the Chan Zuckerberg Initiative AI/ML team as Research Scientist.
- [October 2023] I am presenting a poster on Moscot at Single Cell Genomics 2023 in Engelberg, Switzerland.
- [July 2023] I am presenting a poster on Moscot and SpatialData at the Human Cell Atlas General Meeting 2023, in Toronto.
- [June 2023] I am joining the BioML team, at Microsoft Research New England in Boston, for 3 months.
- [February 2023] Talk at the Statistical Methods for Post Genomic Data workshop, Ghent, 2022.
- [December 2022] Contributed talk at NeurIPS 2022 - LMRL workshop - Modeling Single-Cell Dynamics Using Unbalanced Parameterized Monge Maps.
- [September 2022] Talk at the ECCB 2022 - Workshop on Spatial Transcriptomics and cell-cell communication modelling.
- [August 2022] Contributed to workshop on Spatial Omics Data Analysis 2022.
- [July 2022] Poster at ICML 2022 - Workshop on Computational Biology - Uncertainty Quantification for Atlas-Level Cell Type Transfer.
- [May 2022] Organized virtual workshop Spatial transcriptomics data analysis in Python Single Cell Omics Germany.
- [April 2022] Instructor at the Swiss Institute for Bioinformatics course on Advanced topics in single-cell analysis in Basel.